Software
The code is written in C++ and uses the Object-Oriented Programming (OOP) paradigm. Class templates are used in order to simplify the extension of the 2D model to the 3D space. The source code is documented using Doxygen. The project is accessible on the GitLab repository.
Gitlab EpiCells Project
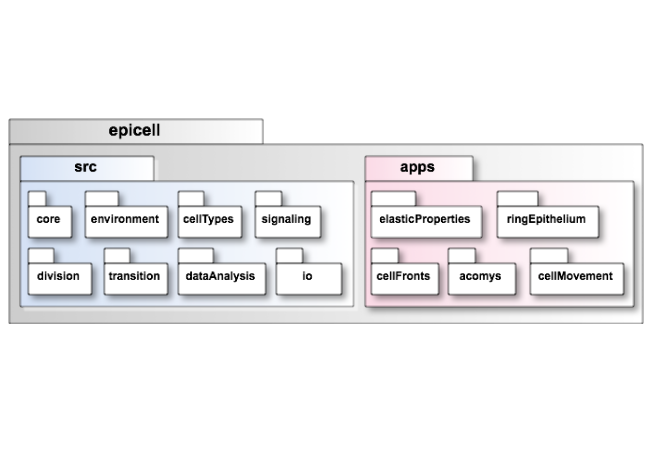
The source code and the application codes are separated in different packages, src and apps, respectively.
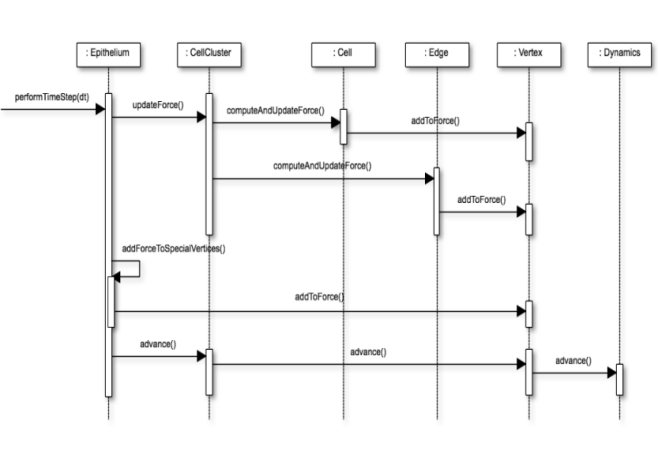
The core component in src includes classes allowing to describe a cell monolayer, including single cells and their neighbourhood relationship. It also allows to define the dynamics of the model driving the movement of the cell vertices over time.
Additional components allow the proliferation of cells, the application of external forces, the occurrence of topological transitions,
the management of different cell types and the cell signalling.